Thanks for the Band Diagram Tutorial for Quantum ESPRESSO
Dependence
-
Quantum Espresso
-
Python3 + Numpy + Matplotlib
Workflow of band diagram
The procedures in this workflow should use the same outdir and prefix configurations.
pw.x
&control and etc. settings for each procedure:
vc-relax
1
2
&control
calculation='vc-relax',
scf
1
2
&control
calculation='scf',
nscf
1
2
&control
calculation='nscf',
bands
1
2
&control
calculation='bands',
after CELL_PARAMETERS section, involve
1
2
3
4
5
K_POINTS crystal
k_points_in_total
a b c wk
a b c wk
a b c wk
The K_POINTS here should follow certain K-path.
The k points settings share the same file coord.txt as in How to Generate QHA Input.
bands.x
The input for bands.x would be like
1
2
3
4
5
&BANDS
prefix="the_prefix"
outdir="the_outdir"
filband="Band.dat"
/
Plot band diagram
Learn about the Quantum ESPRESSO output from bands.x
There are several output types (supposed using filband="Band.dat" in the input for bands.x):
-
Band.dat.gnuwith bands in eV, directly plottable using gnuplot -
Band.dat.rapwith symmetry information, to be read by plotting codeplotband.x` -
Band.datcontaining the band structure, in a format suitable for plotting codeplotband.x -
bandx.out -
scf.outornscf.outProvide the Fermi Energy.
Here, we are using Band.dat to provide band energy, K_Path, and high-symmetry information, scf.out to provide Fermi Energy.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
import numpy as np
import matplotlib.pyplot as plt
import os
import re
def dist(a,b):
# calculate the distance between vector a and b
d = ((a[0]-b[0]) ** 2 + (a[1]-b[1]) ** 2 + (a[2]-b[2]) ** 2)**0.5
return d
def read_fermi(file_name):
# Read the fermi energy in scf.out
fermi = 0
with open(file_name, "r") as f:
lines = f.readlines()
for line in lines:
if "the Fermi energy" in line:
fermi = float(line.split()[4])
return fermi
def read_bnd(file_name):
# Read the bands in Band.dat
coord_regex = r"^\s+(.\d+.\d+)\s+(.\d+.\d+)\s+(.\d+.\d+)$"
x_coord = []
x = []
bands = dict()
with open(file_name, "r") as f:
lines = f.readlines()
for i in range(len(lines)):
line = lines[i]
match = re.match(coord_regex,line)
if match:
x_coord.append([float(match.group(1)), float(match.group(2)), float(match.group(3)) ])
bandddd = lines[i+1] + lines[i+2]
bandddd = bandddd.split()
for j in range(len(bandddd)):
if j not in bands.keys():
bands[j] = []
bands[j].append(float(bandddd[j]))
for i in range(len(x_coord)) :
if i == 0:
x.append(0)
else:
x.append( x[-1] + dist(x_coord[i], x_coord[i-1]))
return bands,x
def plot(bands, x, fermi):
xaxis = [min(x),max(x)]
for i in bands.values():
plt.plot(x, i, color=color_dic[pressure], lw=0.2)
plt.plot(xaxis, [fermi, fermi], color="#66ccff",ls="solid", alpha = 0.5,lw = 1.2)
plt.xlim(xaxis)
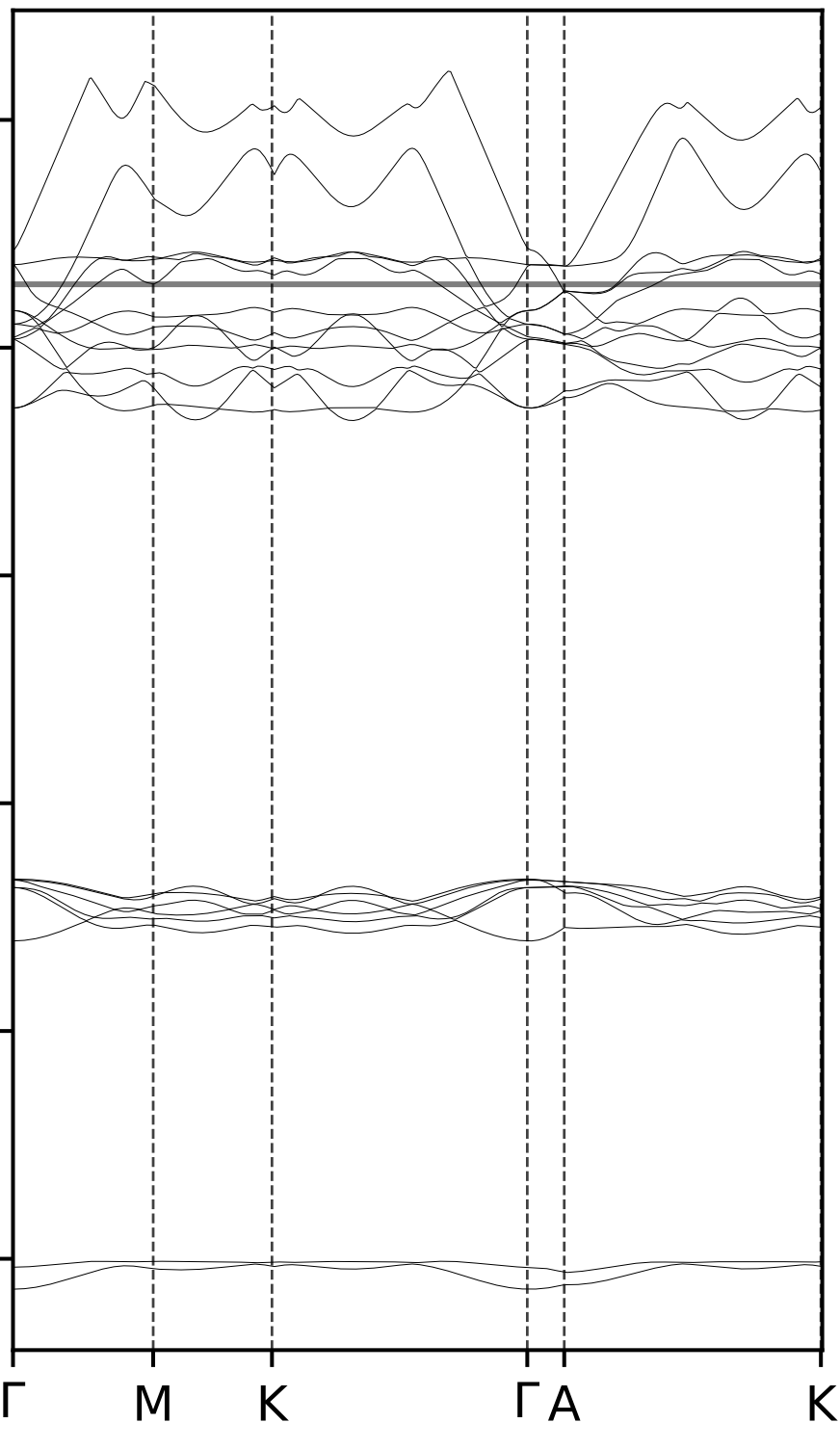
- Plot example: